
GOALS

Electrical Patterning of the Heart
On the tissue/organ scale, the heart is one of the most rapid examples of autonomous, rhythmic, electrochemical oscillation in biology. Our group investigates core questions related to the developmental emergence and optimization of bioelectric signal generation and propagation.

Cell Fate Determination
Embryological development involves the progressive maturation of progenitor cells towards a terminally differentiated state. The long-term goals of our laboratory seek to understand the cellular, molecular, and biophysical events that dictate how cardiovascular cell lineages are established.

Cellular Integration/Organogenesis
Formation of multi-cellular organ systems requires the integration of divergent cell types. Using direct in vivo cell tracing techniques, we focus on identifying the developmental ontogeny and the molecular mechansims that integrate the various cellular components of the heart's electrical system.

RESEARCH
Current research projects in our laboratory are focused on the developmental emergence of cardiac biorythmicity. Effective pumping of blood depends on the rhythmic beating of cardiac muscle. In the mature heart, the timing and coordination of contraction is controlled by a specialized collection of cells referred to as the heart's conduction system. Development of this conduction system entails the de novo formation of a biorhythmic cellular circuit within a rudimentary, albeit functional, embryonic heart. The process of cardiac electrical maturation, therefore, provides a unique model system which informs core biological concepts related to the emergence and optimization of rhythmic electrochemical signal generation, as well as providing insight into the pathophysiology of human arrhythmic disorders. We are currently utilizing classical experimental embryological techniques in combination with state-of-the-art somatic transgenesis and cellular imaging to address: i) the developmental mechanisms that specify autonomous rhythmic signal generation, ii) the processes that allow for effective downstream transmission of these signals, and iii) the morphogenetic patterning events that align the various sub-components of the heart leading to proper spatiotemporal coordination of the cardiac cycle.
Right - High-speed imaging of electrical impulse propagation through a section of embryonic cardiac tissue.
DEVELOPMENTAL PHYSIOLOGY
Modern imaging modalities are allowing for a wide variety of previously inaccessible topics to now be explored in developing tissues. Our group is actively adapting and applying new technologies for labeling and imaging cells in the developing heart in order to gain novel insights into cardiac physiological maturation at subcellular resolution.
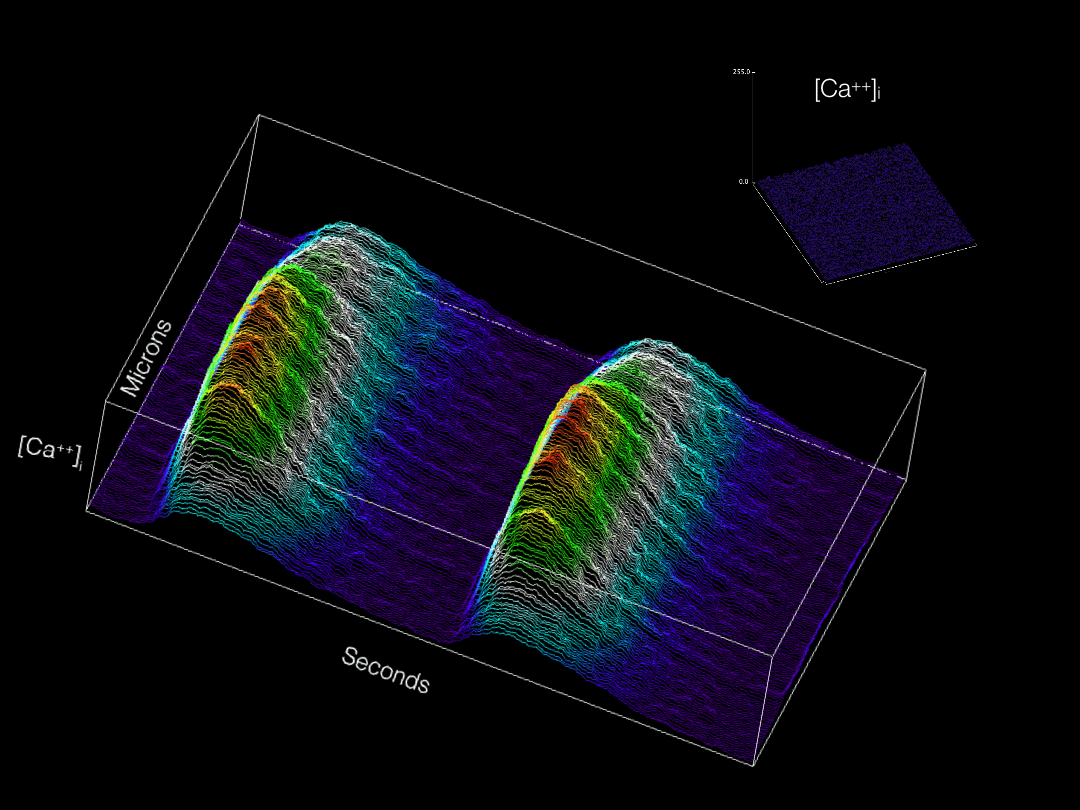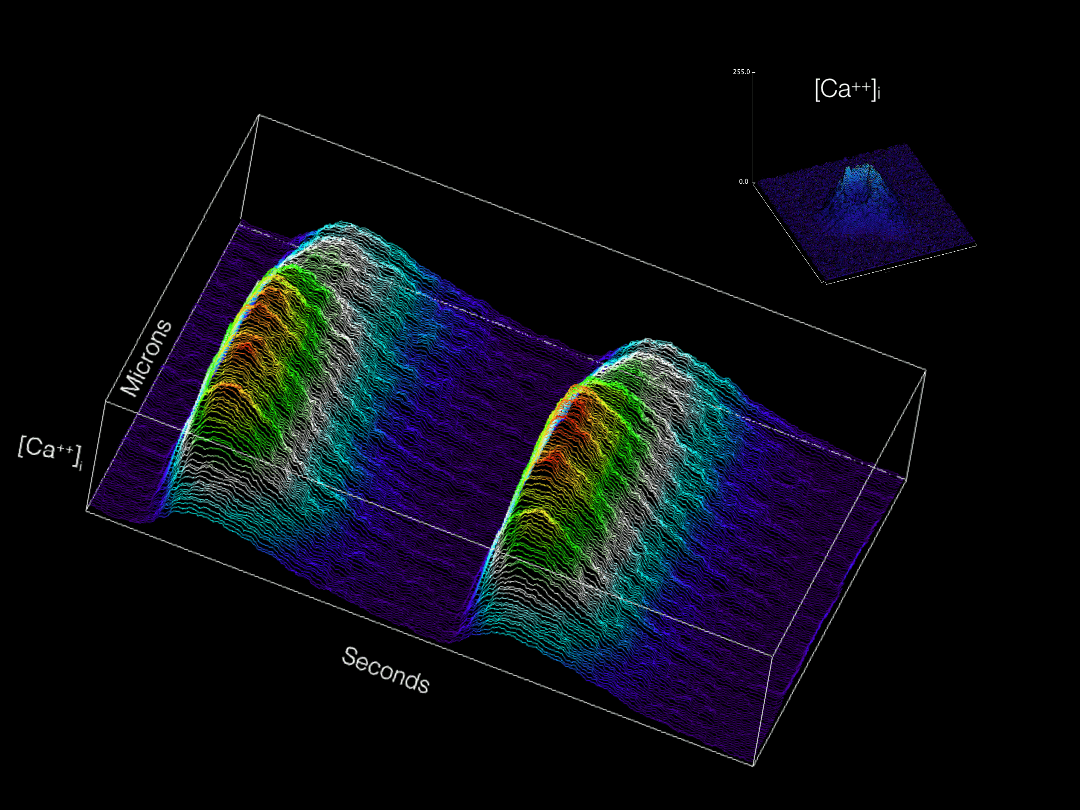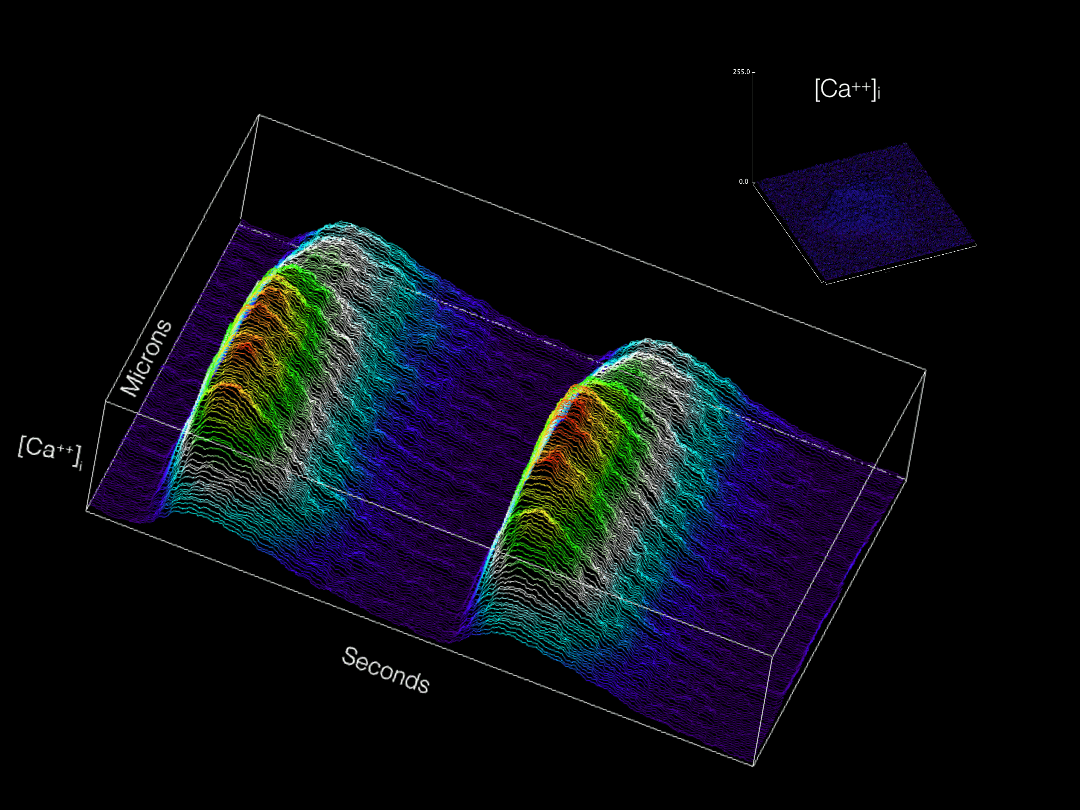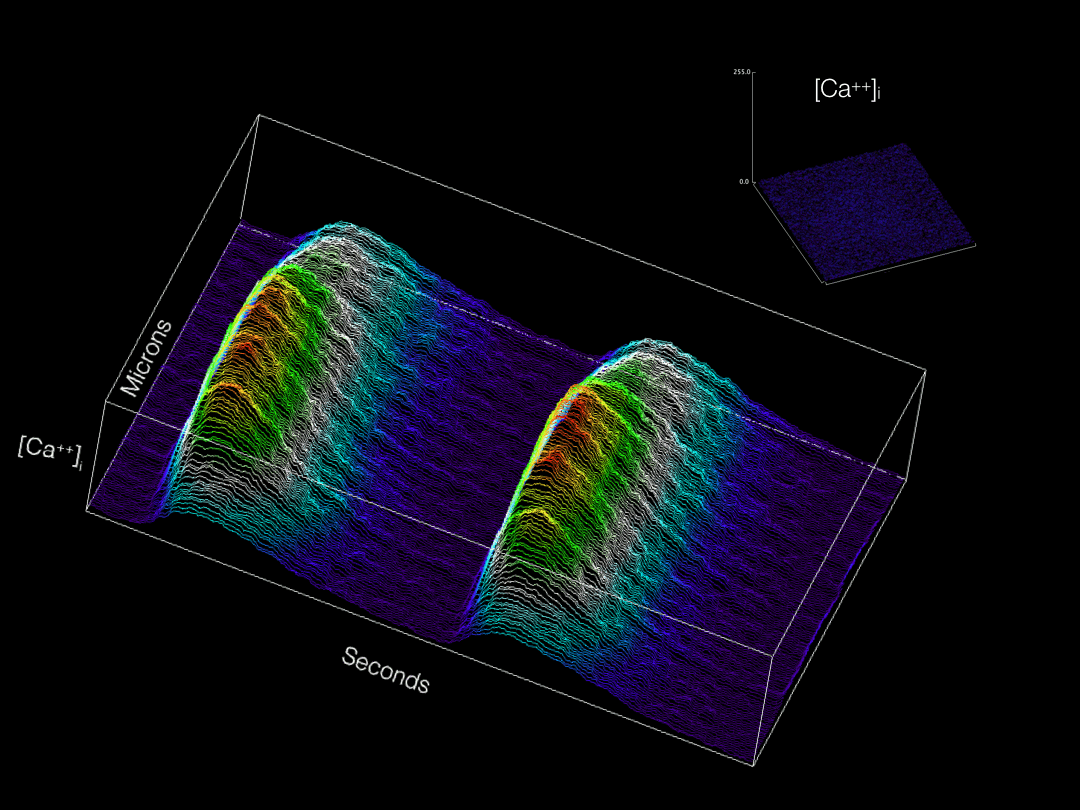
Calcium transients optically recorded from an embryonic cardiomyocyte
PUBLICATIONS
1.
2.
3.
4.
5.
6.
7.
8.
9.
10.
11.
12.
13.
14.
15.
16.
17.
18.
19.
20.
21.
22.
23.
24.
Giesbrecht K, Rossi S, Liu S, Mukherjee S, Bressan M, Griffith BE (2025) An anatomically informed computational fluid dynamics modeling approach for quantifying hemodynamics in the developing heart. PLoS One 20(5):
Henley, T., Goudy, J., Easterling, M., Donley, C., Wirka, R., & Bressan, M. (2023). Local tissue mechanics control cardiac pacemaker cell embryonic patterning. Life Science Alliance, 6(6).
Thomas, K., Henley, T., Rossi, S., Costello, M.J., Polacheck, W., Griffith, B.E., and Bressan, M. (2021). Adherens junction engagement regulates functional patterning of the cardiac pacemaker cell lineage. Developmental Cell.
Easterling, M., Rossi, S., Mazzella, A.J., and Bressan, M. (2021). Assembly of the Cardiac Pacemaking Complex: Electrogenic Principles of Sinoatrial Node Morphogenesis. Journal of Cardiovascular Development and Disease 8, 40.
Wang L., Ma H., Huang P., Xie Y., Near D., Wang H., Xu J., Yang Y., Xu Y., Garbutt T., Zhou Y., Liu Z., Yin C., Bressan M., Taylor J.M., Liu J. and Qian L. (2020) Downregulation of Beclin1 promotes direct cardiac reprogramming. Sci Transl Med.
Guzzetta A, Koska M, Rowton M, Sullivan KR, Jacobs-Li J, Kweon J, Hidalgo H, Eckart H, Hoffmann AD, Back R, Lozano S, Moon AM, Basu A, Bressan M, Pott S, Moskowitz IP. Hedgehog-FGF signaling axis patterns anterior mesoderm during gastrulation. Proc Natl Acad Sci U S A. 2020 Jun 19;. doi: 10.1073/pnas.1914167117. PubMed PMID: 32561646.
Goudy J, Henley T, Méndez HG, Bressan M. Simplified platform for mosaic in vivo analysis of cellular maturation in the developing heart. Sci Rep. 2019 Jul 24;9(1):10716. doi: 10.1038/s41598-019-47009-7. PubMed PMID: 31341189; PubMed Central PMCID: PMC6656758.
Henley, T., Thomas, K., Bressan, M. Microinjection-based System for In Vivo Implantation of Embryonic Cardiomyocytes in the Avian Embryo. J. Vis. Exp. (144), e59267, doi:10.3791/59267 (2019).
Bressan M, Henley T, Louie JD, Liu G, Christodoulou D, Bai X, Taylor J, Seidman CE, Seidman J, Mikawa T. Dynamic cellular integration drives functional assembly of the heart's pacemaker complex. Cell Reports. 2018 May 22;23(8):2283-91.
Thomas K, Goudy J, Henley T, Bressan M. Optical Electrophysiology in the Developing Heart. J Cardiovasc Dev Dis. 2018 May 11;5(2). pii: E28. doi: 10.3390/jcdd5020028. Review. PubMed PMID: 29751595.
Kuyumcu-Martinez MN, Bressan MC. Rebuilding a broken heart: lessons from developmental and regenerative biology. Development. 2016 Nov 1;143(21):3866-3870. Review. PubMed PMID: 27803055.
Bressan M, Liu G, Louie JD, Mikawa T. Cardiac Pacemaker Development from a Tertiary Heart Field. Etiology and Morphogenesis. Chapter 39. 2016; PMID:29787147.
Bressan M, Mikawa T. Avians as a model system of vascular development. Methods Mol Biol. 2015;1214:225-42. doi: 10.1007/978-1-4939-1462-3_14. PubMed PMID: 25468608; PubMed Central PMCID: PMC4582444.
Bressan M, Louie JD, Mikawa T. Hemodynamic forces regulate developmental patterning of atrial conduction. PLoS One. 2014; 9(12):e115207.
Bressan M, Yang PB, Louie JD, Navetta AN, Garriock RJ, Mikawa T. Reciprocal myocardial-endocardial interactions pattern atrioventricular junction conduction delay. Development. 2014 Oct; 141: 4149-4157.
Hua LL, Vedantham V, Barnes RM, Hu J, Robinson AS, Bressan M, Srivastava D, Black BL. Specification of the mouse cardiac conduction system in the absence of Endothelin signaling. Dev Biol. 2014 Sep 15; 393(2):245-54.
Bressan M, Liu G, Mikawa T. Early mesodermal cues assign avian cardiac pacemaker fate potential in a tertiary heart field. Science. 2013 May 10; 340(6133): 744-8.
Maya-Ramos L, Cleland J, Bressan M, Mikawa T. Induction of the Proepicardium. J. Dev. Biol. 2013, 1(2): 82-91
Liu J, Bressan M, Hassel D, Huisken J, Staudt D, Kikuchi K, Poss KD, Mikawa T, Stainier DY. A dual role for ErbB2 signaling in cardiac trabeculation. Development. 2010 Nov; 137(22):3867-75.
Bressan M, Davis P, Timmer J, Herzlinger D, Mikawa T. Notochord-derived BMP antagonists inhibit endothelial cell generation and network formation. Dev Biol. 2009 Feb 1; 326(1):101-11.
Guillaume, R., Bressan, M., Hezlinger, D. Paraxial mesoderm contributes stromal cells to the developing kidney. Dev Biol. 2009 May; 329(2): 169-175.
Pallante BA, Duignan I, Okin D, Chin A, Bressan MC, Mikawa T, Edelberg JM. Bone marrow Oct3/4+ cells differentiate into cardiac myocytes via age-dependent paracrine mechanisms. Circ Res. 2007 Jan 5; 100(1):e1-11.
McDermott DA, Bressan MC, He J, Lee JS, Aftimos S, Brueckner M, Gilbert F, Graham GE, Hannibal MC, Innis JW, Pierpont ME, Raas-Rothschild A, Shanske AL, Smith WE, Spencer RH, St John-Sutton MG, van Maldergem L, Waggoner DJ, Weber M, Basson CT. TBX5 genetic testing validates strict clinical criteria for Holt-Oram syndrome. Pediatr Res. 2005 Nov; 58(5):981-6.
Veugelers M, Bressan M, McDermott DA, Weremowicz S, Morton CC, Mabry CC, Lefaivre JF, Zunamon A, Destree A, Chaudron JM, Basson CT. Mutation of perinatal myosin heavy chain associated with a Carney complex variant. N Engl J Med. 2004 Jul 29; 351(5):460-9.
CURRENT LAB MEMBERS






Michael Bressan PhD
Principle Investigator
Assistant Professor
Department of Cell Biology and Physiology
McAllister Heart Insititute
Training:
Postdoctoral Fellow
University of California, San Francisco
Cardiovascular Research Institute
PhD
Weill Medical College of Cornell University
Dept. of Physiology, Biophysics, and Systems Biology
B.S.
Lafayette College
Dept. of Biology
Kathryn Scherrer MS
Lab Manager/Research Technician
Training:
M.S.
Colorado School of Mines
Dept. of Quantitative Biosciences and Engineering
B.S.
Colorado School of Mines
Dept. of Biochemical Engineering
Kirsten Giesbrecht PhD
Postdoctoral Fellow
Training:
PhD
University of North Carolina - Chapel Hill
Dept. of Math
B.S.
Centre College
Dept. of Math
Ashlyn Laidman
Graduate Student - 3rd Year
Training:
B.S.
Texas Christian University
Dept. of Biology and Dept. of Chemistry
Trevor Henley
Graduate Student - 2nd Year
Training:
B.S.
University of North Carolina - Chapel Hill
Dept. of Biology
Kaitlyn Huizar
Graduate Student - 2nd Year
Training:
B.S.
Angelo State University
Dept. of Psychology
PREVIOUS LAB MEMBERS
Trevor Henley
-
Laboratory Manager 2016-2023
-
B.S. in Biology (University of North Carolina - Chapel Hill)
-
Currently a graduate student at UNC CH
Julie Goudy
-
Research Technician 2016-2022
-
M.S. in Physiology and Developmental Biology (Brigham Young University)
-
B.S. in Physiology and Developmental Biology (Brigham Young University)
-
Currently Clinical Research Specialist at UNC
Kandace Thomas
-
Graduate Student 2016-2021
-
B.S. in Biology (Eastern University)
-
Currently Assistant Professor at EU
Simone Rossi PhD
-
Visiting postdoctoral fellow 2016-2022
-
PhD in mathematics (IST-EPFL)
-
MS/BS in physics (Università degli Studi di Milano)
-
Currently Software Engineer at Align Technology
Marietta Easterling PhD
-
Postdoctoral Fellow 2019-2022
-
PhD in Biology (Washington State University)
-
BS in Biology (College of Charleston)
-
Currently Field Application Scientist at 10x Genomics
RECENT ROTATION STUDENTS
-
Kaitlyn Huizar Summer 2025
-
Katherine Long Winter 2024-2025
-
Trevor Henley Fall 2024
-
Ashlyn Laidman Winter 2023-2024
-
Anh Luu Winter 2023-2024
FURRY LAB MEMBERS

Juno and Copper

Cami and Zwitter

Ruthie
Lab Outings and Activities


Department Pumpkin Carving 2024

Lab Lunch Fall Rotation 2024

Cytoskeletal Conference 2024
CONTACT
Employment Inquiries:
The laboratory is always looking for motivated postdocs and students. Please send a CV and a description of research interests to Michael Bressan.
Currently accepting rotation students!
(As of June 2025)
Office
University of North Carolina, Chapel Hill
111 Mason Farm Rd. MBRB 6341c
Chapel Hill, NC
27599
Tel: 919.843.9455
Laboratory
111 Mason Farm Rd. MBRB 6335
Chapel Hill, NC
27599
Tel: 919.962.3008
